Professor Yang
 Know more>>
Professor
Department of Chemical Biology, Xiamen University
Know more>>
Professor
Department of Chemical Biology, Xiamen University
Contact Information
Yang's LaboratoryRoom 532, Lujiaxi Building, College of Chemistry and Chemical Engineering, Xiamen University, Xiamen 361005, China
Ph: +86 (0) 592-218 7601cyyang@xmu.edu.cn
Yahong Chen's paper accepted by Nat Protoc
2023-10-05 09:56:00

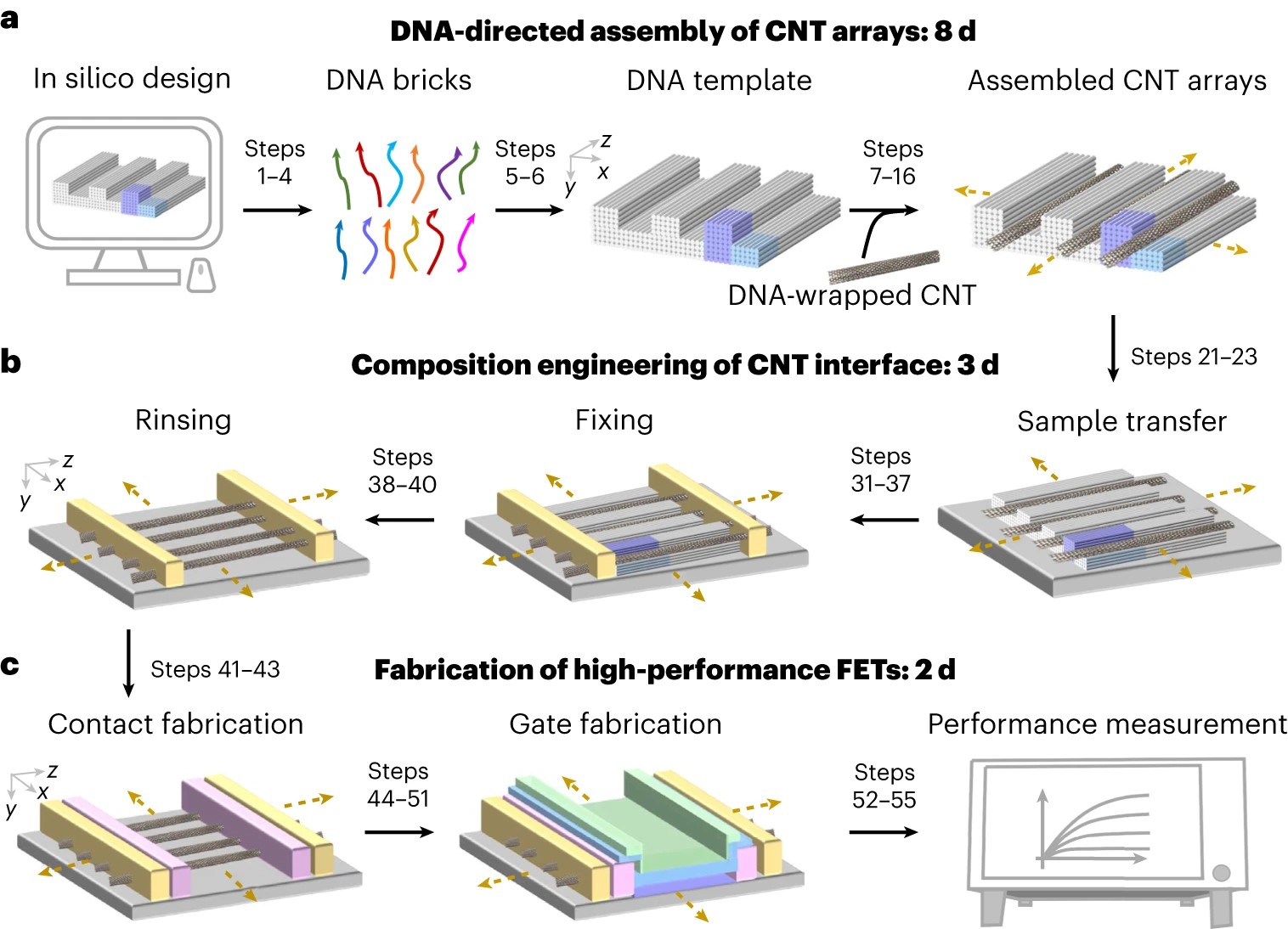
Structural DNA nanotechnology can be programmed into complex designer structures with molecular precision for directing a wide range of inorganic and biological materials. However, the use of DNA-templated approaches for the fabrication and performance requirements of ultra-scaled semiconductor electronics is limited by its assembly disorder and destructive interface composition. In this protocol, using carbon nanotubes (CNTs) as model semiconductors, we provide a stepwise process to build ultra-scaled, high-performance field-effect transistors (FETs) from micron-scale three-dimensional DNA templates. We apply the approach to assemble CNT arrays with uniform pitches scaled between 24.1 and 10.4 nm with yields of more than 95%, which exceeds the resolution limits of conventional lithography. To achieve highly clean CNT interfaces, we detail a rinsing-after-fixing step to remove residual DNA template and salt contaminations present around the contact and the channel regions, without modifying the alignment of the CNT arrays. The DNA-templated CNT FETs display both high on-state current (4–15 μA per CNT) and small subthreshold swing (60–100 mV per decade), which are superior to previous examples of biotemplated electronics and match the performance metrics of high-performance, silicon-based electronics. The scalable assembly of defect-free three-dimensional DNA templates requires 1 week and the CNT arrays can be synthesized within half a day. The interface engineering requires 1–2 d, while the fabrication of high-performance FET and logic gate circuits requires 2–4 d. The structural and performance characterizations of molecular-precise DNA self-assembly and high-performance electronics requires 1–2 d. The protocol is suited for users with expertise in DNA nanotechnology and semiconductor electronics.