Professor Yang
 Know more>>
Professor
Department of Chemical Biology, Xiamen University
Know more>>
Professor
Department of Chemical Biology, Xiamen University
Contact Information
Yang's LaboratoryRoom 532, Lujiaxi Building, College of Chemistry and Chemical Engineering, Xiamen University, Xiamen 361005, China
Ph: +86 (0) 592-218 7601cyyang@xmu.edu.cn
Mingyin Li's paper accepted by ACS Appl Mater Interfaces
2024-10-22 17:04:46

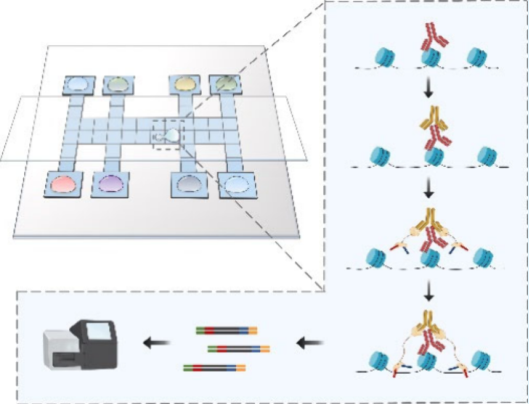
Mapping genome-wide DNA-protein interactions (DPIs) provides insights into the epigenetic landscape of complex biological systems and elucidates the mechanisms of epigenetic regulation in biological progress. However, current technologies in DPI profiling still suffer from high cell demands, low detection sensitivity, and large reagent consumption. To address these problems, we developed DMF-ChIP-seq that builds on digital microfluidic (DMF) technology to profile genome-wide DPIs in a highly efficient, cost-effective, and user-friendly way. The entire workflow including cell pretreatment, antibody recognition, pA-Tn5 tagmentation, fragment enrichment, and PCR amplification is programmatically manipulated on a single chip. Leveraging closed submicroliter reaction volumes and a superhydrophobic interface, DMF-ChIP-seq presented higher sensitivity in peak enrichment than other current methods, with high accuracy (Pearson Correlation Coefficient (PCC) > 0.86) and high repeatability (PCC > 0.92). Furthermore, DMF-ChIP-seq was capable of processing the samples with as few as 8 cells while maintaining a high signal-to-noise ratio. By applying DMF-ChIP-seq, H3K27ac histone modification of early embryonic cells during differentiation was profiled for the investigation of epigenomic landscape dynamics. With the benefits of high efficiency and sensitivity in DPI analysis, the system provides great promise in studying epigenetic regulation during various biological processes.