Professor Yang
 Know more>>
Professor
Department of Chemical Biology, Xiamen University
Know more>>
Professor
Department of Chemical Biology, Xiamen University
Contact Information
Yang's LaboratoryRoom 532, Lujiaxi Building, College of Chemistry and Chemical Engineering, Xiamen University, Xiamen 361005, China
Ph: +86 (0) 592-218 7601cyyang@xmu.edu.cn
Xing Xu's paper accepted by Sci China Chem
2024-04-16 16:54:33

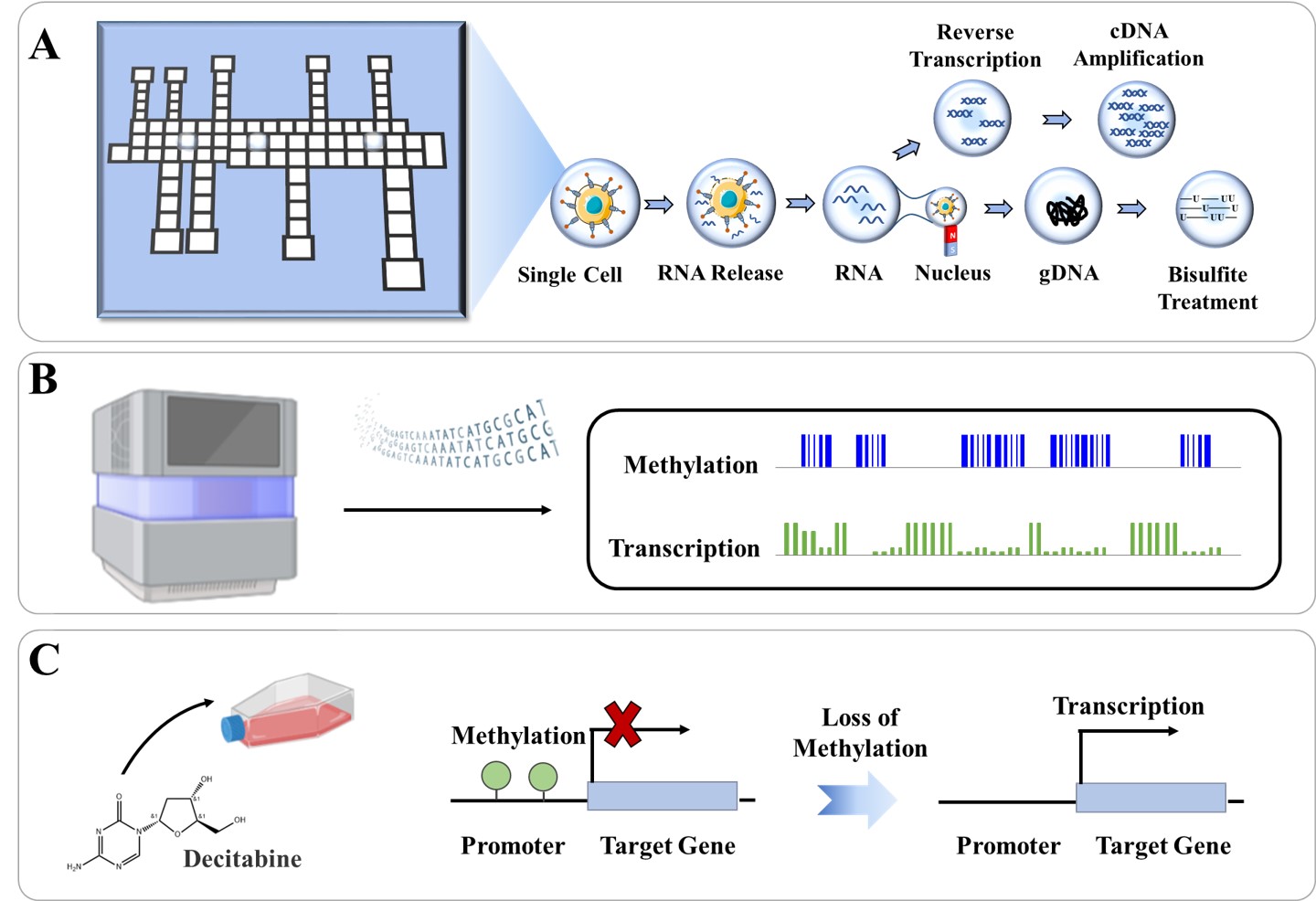
Single-cell joint analysis of methylome and transcriptome reveals how the methylation regulates the transcriptional activity. However, traditional bench-top protocols for single-cell DNA methylation and RNA transcription co-detection are labor-intensive, cost-ineffective and contaminant-prone. Herein we establish DMF-scMT-seq, a highly-efficient and cost-effective method to simultaneously analyze single-cell DNA methylation and transcriptional activity based on digital microfluidics. DMF-scMT-seq automates the workflow of single-cell isolation, cellular hypotonic lysis, nucleic acid separation and methylome/transcriptome library construction in a contactless and addressable way. The system ensures high accuracy (R>0.85), high gene detection ability (14697 genes per cell at 4 million sequencing depth), and high CpG coverage (677198 CpG sites per cell at 1 million sequencing depth). By using DMF-scMT-seq, the relationship of DNA methylation and RNA transcription under different genomic contexts is resolved. We further apply DMF-scMT-seq to study the dynamics of transcription regulation with methylation-inhibiting anti-tumor Decitabine, and identify the methylated promoter/gene body driven genes in response to Decitabine treatment for the first time. DMF-scMT-seq facilitates the construction of the correlation of DNA methylation and transcriptional activity at the single-cell level in a flexible, sensitive and accurate way, which is anticipated to be a powerful tool in studying single-cell biological systems.